What is ExpreSEd?
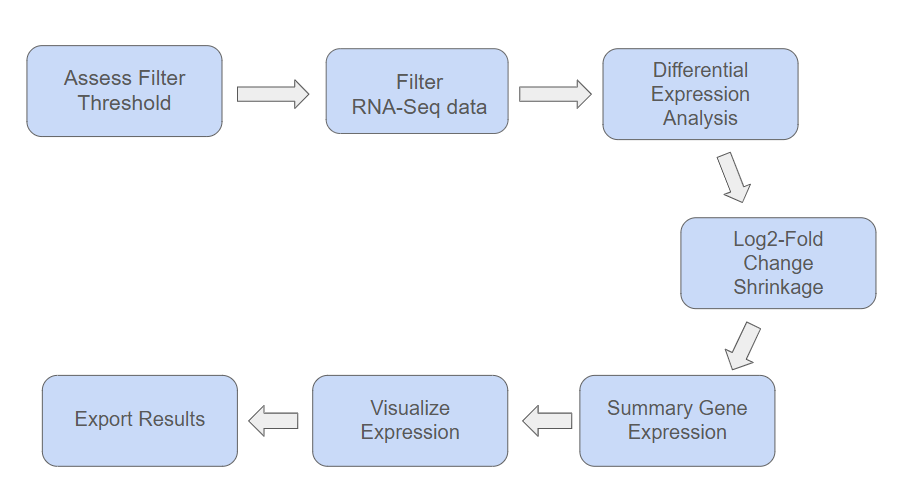
ExpreSEd is a package that provides a streamlined pipeline for differential expression analysis using SummarizedExperiment objects, and raw counts data and sample metadata files. Input bulk RNA-seq data, and ExpreSEd will output an optimal filter threshold matrix, log2-fold change shrunken differential analysis results, a summary of expression patterns, and a volcano plot. ExpreSEd is compatible with R, command line, Docker, and Nextflow programs.
Package Components
├── DESCRIPTION
├── ExpreSEd.Rproj
├── LICENSE
├── LICENSE.md
├── NAMESPACE
├── R
│ ├── DESeq2_analysis.R
│ ├── ExpreSEd-package.R
│ ├── data-documentation.R
│ ├── determine_filter_threshold.R
│ ├── export.R
│ ├── filter_low_exp_gene.R
│ ├── shrinkage.R
│ ├── summarize_genes.R
│ └── volcano_plot.R
├── README.Rmd
├── README.md
├── _pkgdown.yml
├── data
│ └── example_se.rda
├── data-raw
│ └── example_se.R
├── exec
│ └── ADS8192.R
├── man
│ ├── ExpreSEd-package.Rd
│ ├── determine_filter_threshold.Rd
│ ├── example_se.Rd
│ ├── export_outputs.Rd
│ ├── filter_low_exp_genes.Rd
│ ├── gene_regulation_summary.Rd
│ ├── generate_volcano.Rd
│ ├── log2_shrinkage.Rd
│ └── run_DESeq2.Rd
├── tests
│ ├── testthat
│ │ ├── test-DESeq2_analysis.R
│ │ ├── test-determine_filter_threshold.R
│ │ ├── test-export.R
│ │ ├── test-filter_low_exp_genes.R
│ │ ├── test-shrinkage.R
│ │ ├── test-summarize_genes.R
│ │ └── test-volcano_plot.R
│ └── testthat.R
└── vignettes
├── about.Rmd
├── cli-analysis.Rmd
├── cli-analysis.html
├── getting-started.Rmd
├── prebuilt_data.RData
├── r-analysis.Rmd
├── r-analysis.html
└── workflow.png